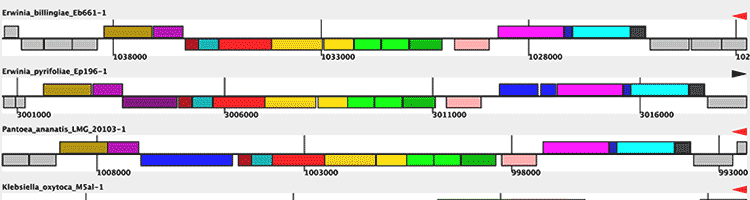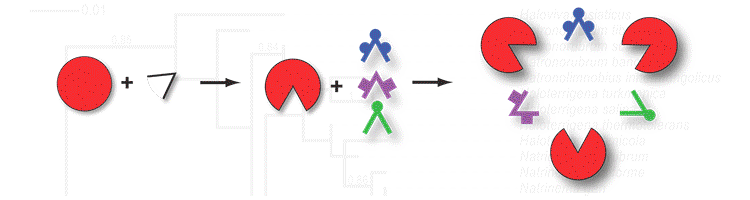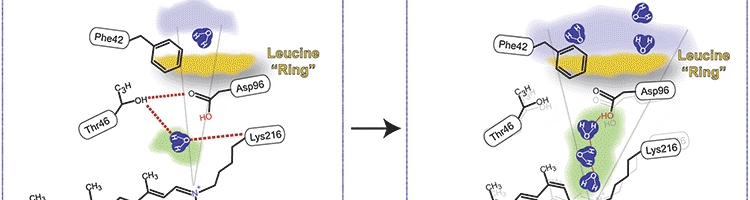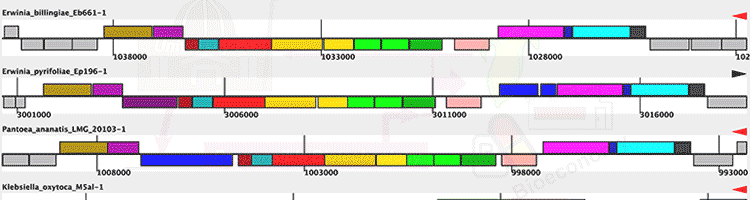
 Photo Credits: BMC Bioinformatics, PLoS One, Cell Press, ACS Synthetic Biology |
SEPTEMBER 2021 – A WEB PAGE UPDATE IS IN PROGRESS.
UPDATES ON TISSUE ENGINEERING AND EDUCATIONAL INITIATIVES ARE COMING
Microbial GenomicsDNA sequencing technologies are providing incredible opportunities for discovery and understanding in the biological sciences. This is particularly true in the microbial sciences, where new tools now offer views into the genomes of many model and non-model microorganisms and into the metagenomic composition and dynamics of microbial communities. However, studies interpreting the functional expression of these genomes (what the genomes do and how they do it) are lagging far behind. We are interested developing experimental and computational approaches to bridge this knowledge gap. Systems BiologyOne of the major goals of Systems Biology is to decipher the structures and understand the dynamics of cellular networks in a way that allows us to predicatively rewire these cellular circuits for our own purpose. This ability will facilitate the rewiring of disease-perturbed networks back to health and the engineering of novel cellular factories. We are interested in utilizing the tools and frameworks of Systems Biology to learn more about how structure and dynamics in gene regulatory networks lead to the emergence of complex phenotypes. Synthetic BiologyDeveloping the means to efficiently engineer biological systems has broad application, from fighting disease to engineering microbial factories and beyond. We are interested in pursuing questions related to standards development, chassis/circuit interactions, and the interface between genomic systems and synthetic biology. Molecular BiophysicsThe chemical and biophysical properties of macromolecules are at the core of most biological phenotypes. Developing a molecular-level understanding of key macromolecules is therefore critical to both understanding and engineering more complex biological systems. When possible, we work with collaborators to develop a better biophysical understanding of macromolecules that may be of further scientific and/or biotechnological interest. To date, studies have focused primarily on the haloarchaeal opsins and haloarchaeal lipid systems. |
